 With this menu entry, the user can map a single DNA sequence or collection of DNA fragments to any other DNA sample in the library or genome. Another great feature is the menu entry "Serial Cloning Window". For example, there is a "Genetic Sequence Window" that contains an extensive list of genetic sequences that this software can automatically map and analyze. over-expressing DNF to see whether this would cause the plants to be. This is just a small sampling of the features that this great software program has to offer.Īnother great feature of this product is that users can manually select which windows and menus of the utility application they want to use to perform their tasks. mutant carrying the gi mutation combined with mutations in the four CDF genes (CDF. The software has a large variety of useful functions such as real-time generation of mapped fragments, correction of errors in sequences, identification of individual genes, multipleogenicity detection, hybridization detection, population genetic analysis, scoring methods, and database support. 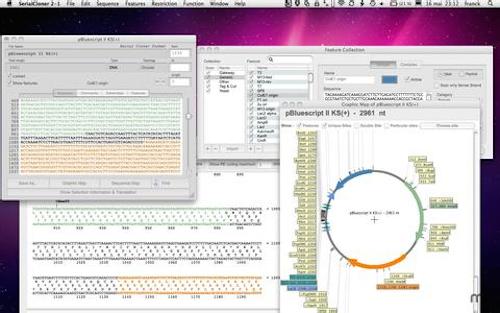
Possible outcomes of these observations are discussed. This method called TDNAscan was developed to be an open source software. T-DNA insertions at the 5 end of the AtTERT gene or in the region upstream of the ATG start codon led to the activation of putative regulatory elements. Our method enables the detection of the corresponding T-DNA insertion orientation and zygosity, as well as insertion annotation. Then we take a standard material and add in a MoGraph Multishader to the colour channel. To start with we clone the flag with a cloner and sort out the arrangement. The software is able to read and write DNA Strider-readable files and automatically import and exports data into the universal FASTA file format. Here, we present a next generation sequencing (NGS)-based platform to facilitate the identification of complete and truncated T-DNA insertions. In this tutorial, we learn how to apply different materials to clones, in this instance, a flag. The software has a graphical user interface (GUI) that makes sequencing and visualizing data easy and fast. All of our incoming plasmids are sequenced at least. This is the most important step in ensuring quality, as it will catch any errors made during the cloning, sample handling, or sequence assembly steps. The manual will show you the basic workflow that is involved with sequencing, cloning, and analyzing DNA samples. Once the physical sample and its associated information are at Addgene, we are ready to sequence verify the plasmids. 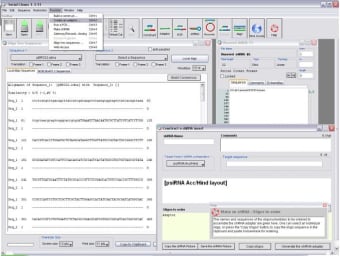
To give you an idea about how Serial Cloner works, let us take a look at the program's manual and how to use it. The software is available for purchase on the web and for download straight from the company's website. This software can be used by anyone with a Windows computer. library construction from a wild individual clone of entire gene PCR with. Serial Cloner is an awesome software tool that allows users of molecular biology research to quickly and easily sequence, analyze, visualize, analyze, and interpret the data obtained during their experiments. gene G mutated and tagged ( tagging : insertion of transposon or T - DNA in G ). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
